Assemblies
Assembly Name:
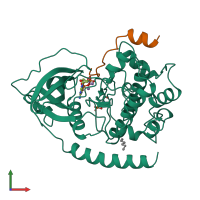
cAMP-dependent protein kinase inhibitor alpha and cAMP-dependent protein kinase catalytic subunit alpha
Multimeric state:
hetero dimer
Accessible surface area:
15935.12 Å2
Buried surface area:
3698.5 Å2
Dissociation area:
1,115.04
Å2
Dissociation energy (ΔGdiss):
7.42
kcal/mol
Dissociation entropy (TΔSdiss):
9.17
kcal/mol
Symmetry number:
1
PDBe Complex ID:
PDB-CPX-153239
Assembly Name:
cAMP-dependent protein kinase inhibitor alpha and cAMP-dependent protein kinase catalytic subunit alpha
Multimeric state:
hetero dimer
Accessible surface area:
15897.21 Å2
Buried surface area:
3711.84 Å2
Dissociation area:
1,115.45
Å2
Dissociation energy (ΔGdiss):
6.5
kcal/mol
Dissociation entropy (TΔSdiss):
9.17
kcal/mol
Symmetry number:
1
PDBe Complex ID:
PDB-CPX-153239
Macromolecules
Chains: A, B
Length: 350 amino acids
Theoretical weight: 40.63 KDa
Source organism: Sus scrofa
UniProt:
Pfam: Protein kinase domain
InterPro:
SCOP: Protein kinases, catalytic subunit
Length: 350 amino acids
Theoretical weight: 40.63 KDa
Source organism: Sus scrofa
UniProt:
- Canonical:
 P36887 (Residues: 2-351; Coverage: 100%)
P36887 (Residues: 2-351; Coverage: 100%)
Pfam: Protein kinase domain
InterPro:
- Protein kinase-like domain superfamily
- cAMP-dependent protein kinase catalytic subunit
- Protein kinase domain
- Protein kinase, ATP binding site
- Serine/threonine-protein kinase, active site
- AGC-kinase, C-terminal
SCOP: Protein kinases, catalytic subunit
Chains: I, J
Length: 20 amino acids
Theoretical weight: 2.23 KDa
Source organism: Homo sapiens
Expression system: Not provided
UniProt:
InterPro: cAMP-dependent protein kinase inhibitor
Length: 20 amino acids
Theoretical weight: 2.23 KDa
Source organism: Homo sapiens
Expression system: Not provided
UniProt:
- Canonical:
 P61926 (Residues: 6-25; Coverage: 26%)
P61926 (Residues: 6-25; Coverage: 26%)
InterPro: cAMP-dependent protein kinase inhibitor