Assemblies
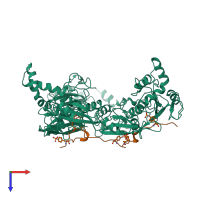
Multimeric state:
hetero hexamer
Accessible surface area:
35894.37 Å2
Buried surface area:
8961.01 Å2
Dissociation area:
1,296.56
Å2
Dissociation energy (ΔGdiss):
-9.71
kcal/mol
Dissociation entropy (TΔSdiss):
26.14
kcal/mol
Symmetry number:
1
PDBe Complex ID:
PDB-CPX-139753
Multimeric state:
hetero hexamer
Accessible surface area:
37634.29 Å2
Buried surface area:
9141.53 Å2
Dissociation area:
1,239.01
Å2
Dissociation energy (ΔGdiss):
-12.04
kcal/mol
Dissociation entropy (TΔSdiss):
26.37
kcal/mol
Symmetry number:
1
PDBe Complex ID:
PDB-CPX-139753
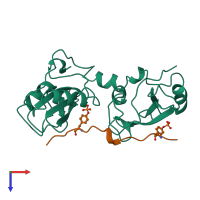
Multimeric state:
hetero dimer
Accessible surface area:
13830.83 Å2
Buried surface area:
2192.42 Å2
Dissociation area:
1,096.21
Å2
Dissociation energy (ΔGdiss):
14.72
kcal/mol
Dissociation entropy (TΔSdiss):
9.46
kcal/mol
Symmetry number:
1
PDBe Complex ID:
PDB-CPX-139750
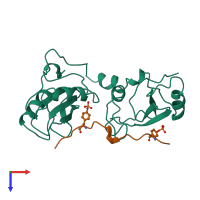
Multimeric state:
hetero dimer
Accessible surface area:
12123.41 Å2
Buried surface area:
2164.43 Å2
Dissociation area:
1,082.22
Å2
Dissociation energy (ΔGdiss):
13.35
kcal/mol
Dissociation entropy (TΔSdiss):
9.54
kcal/mol
Symmetry number:
1
PDBe Complex ID:
PDB-CPX-139750
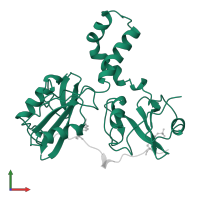
Multimeric state:
hetero dimer
Accessible surface area:
12299.3 Å2
Buried surface area:
2132.6 Å2
Dissociation area:
1,066.3
Å2
Dissociation energy (ΔGdiss):
11.62
kcal/mol
Dissociation entropy (TΔSdiss):
9.45
kcal/mol
Symmetry number:
1
PDBe Complex ID:
PDB-CPX-139750
Multimeric state:
hetero dimer
Accessible surface area:
13944.7 Å2
Buried surface area:
2210.38 Å2
Dissociation area:
1,105.19
Å2
Dissociation energy (ΔGdiss):
11.32
kcal/mol
Dissociation entropy (TΔSdiss):
9.56
kcal/mol
Symmetry number:
1
PDBe Complex ID:
PDB-CPX-139750
Multimeric state:
hetero dimer
Accessible surface area:
14064.78 Å2
Buried surface area:
2070.86 Å2
Dissociation area:
1,035.43
Å2
Dissociation energy (ΔGdiss):
11.09
kcal/mol
Dissociation entropy (TΔSdiss):
9.5
kcal/mol
Symmetry number:
1
PDBe Complex ID:
PDB-CPX-139750
Multimeric state:
hetero dimer
Accessible surface area:
12336.78 Å2
Buried surface area:
2260.72 Å2
Dissociation area:
1,130.36
Å2
Dissociation energy (ΔGdiss):
11.23
kcal/mol
Dissociation entropy (TΔSdiss):
9.55
kcal/mol
Symmetry number:
1
PDBe Complex ID:
PDB-CPX-139750
Macromolecules
Chains: A, C, E, G, I, K
Length: 254 amino acids
Theoretical weight: 28.83 KDa
Source organism: Homo sapiens
Expression system: Escherichia coli
UniProt:
Pfam: SH2 domain
InterPro:
SCOP: SH2 domain
Length: 254 amino acids
Theoretical weight: 28.83 KDa
Source organism: Homo sapiens
Expression system: Escherichia coli
UniProt:
- Canonical:
 P43405 (Residues: 9-262; Coverage: 40%)
P43405 (Residues: 9-262; Coverage: 40%)
Pfam: SH2 domain
InterPro:
- SH2 domain superfamily
- SH2 domain
- SYK/ZAP-70, N-terminal SH2 domain
- Tyrosine-protein kinase SYK/ZAP-70, inter-SH2 domain superfamily
SCOP: SH2 domain
Chains: B, D, F, H, J, L
Length: 18 amino acids
Theoretical weight: 2.28 KDa
Source organism: Homo sapiens
Expression system: Not provided
UniProt:
Pfam: Immunoreceptor tyrosine-based activation motif
InterPro: Phosphorylated immunoreceptor signalling ITAM
Length: 18 amino acids
Theoretical weight: 2.28 KDa
Source organism: Homo sapiens
Expression system: Not provided
UniProt:
- Canonical:
 P07766 (Residues: 186-203; Coverage: 10%)
P07766 (Residues: 186-203; Coverage: 10%)
Pfam: Immunoreceptor tyrosine-based activation motif
InterPro: Phosphorylated immunoreceptor signalling ITAM