EMD-9524
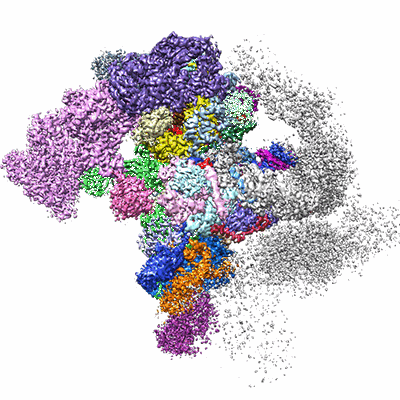
Cryo-EM structure of the activated spliceosome (Bact complex) at 3.5 angstrom resolution
EMD-9524
Single-particle3.5 Å
 Deposition: 18/08/2016
Deposition: 18/08/2016Map released: 21/09/2016
Last modified: 06/11/2019
Name: Bact spliceosomal complex
Summary:
- Complex
- Protein
- Pre-mRNA-splicing factor 8
- Pre-mRNA-splicing helicase BRR2
- Pre-mRNA-splicing factor SNU114
- Pre-mRNA-splicing factor RSE1
- Cold sensitive U2 snRNA suppressor 1
- Pre-mRNA-splicing factor PRP11
- Pre-mRNA-splicing factor RDS3
- RDS3 complex subunit 10
- Pre-mRNA-splicing factor PRP46
- Pre-mRNA-processing protein 45
- Pre-mRNA-splicing factor SLT11
- Pre-mRNA-splicing factor CWC2
- Pre-mRNA-splicing factor CWC15
- Pre-mRNA-splicing factor BUD31
- Pre-mRNA leakage protein 1
- U2 snRNP component IST3
- Pre-mRNA-splicing factor CWC26
- Pre-mRNA-splicing factor ATP-dependent RNA helicase-like protein PRP2
- Pre-mRNA-splicing factor CWC22
- Pre-mRNA-splicing factor CWC24
- Peptidyl-prolyl isomerase CWC27
- Pre-mRNA-splicing factor CEF1
- U2 snRNP component HSH155
- Pre-mRNA-splicing factor CLF1
- Pre-mRNA-splicing factor CWC21
- Pre-mRNA-splicing factor SYF1
- Protein HSH49
- Pre-mRNA-splicing factor SYF2
- Pre-mRNA-processing factor 19
- Pre-mRNA-splicing factor SNT309
- Pre-mRNA-processing factor 17
- Small nuclear ribonucleoprotein-associated protein B
- Small nuclear ribonucleoprotein E
- Small nuclear ribonucleoprotein F
- Small nuclear ribonucleoprotein G
- Small nuclear ribonucleoprotein Sm D3
- Small nuclear ribonucleoprotein Sm D1
- Small nuclear ribonucleoprotein Sm D2
- RNA
- Saccharomyces cerevisiae strain CDRDR_sf_H chromosome VII sequence
- Saccharomyces cerevisiae strain T.52_2H chromosome XII sequence
- U2 snRNA
- Pre-mRNA
- Pre-mRNA
- Ligand
Name: Bact spliceosomal complex
Natural source [1]
Organism: Saccharomyces cerevisiae S288c
Molecular weight
| Experimental | Theoretical | Method |
|---|---|---|
| - | 2 MDa | - |
Complex Portal [1]
| ID | Name | Overlap |
|---|---|---|
| CPX-1651 | PRP19-associated complex | 0.58 |
Name: Pre-mRNA-splicing factor 8
Number of copies: 1
UniProtKB PDBe-KB AlphaFold DB
Number of copies: 1
UniProtKB PDBe-KB AlphaFold DB
Domains
Gene Ontology [20]
Biological process:
spliceosomal snRNP assembly mRNA splicing, via spliceosome generation of catalytic spliceosome for second transesterification step mRNA 3'-splice site recognition spliceosomal tri-snRNP complex assembly mRNA 5'-splice site recognition
Molecular function:
mRNA binding metallopeptidase activity isopeptidase activity U6 snRNA binding pre-mRNA intronic binding U5 snRNA binding U2 snRNA binding U1 snRNA binding
Cellular location:
spliceosomal complex cytoplasm U5 snRNP U4/U6 x U5 tri-snRNP complex nucleus catalytic step 2 spliceosome
spliceosomal snRNP assembly mRNA splicing, via spliceosome generation of catalytic spliceosome for second transesterification step mRNA 3'-splice site recognition spliceosomal tri-snRNP complex assembly mRNA 5'-splice site recognition
Molecular function:
mRNA binding metallopeptidase activity isopeptidase activity U6 snRNA binding pre-mRNA intronic binding U5 snRNA binding U2 snRNA binding U1 snRNA binding
Cellular location:
spliceosomal complex cytoplasm U5 snRNP U4/U6 x U5 tri-snRNP complex nucleus catalytic step 2 spliceosome
Natural source
Molecular weight
| Experimental | Theoretical | Method |
|---|---|---|
| - | 265 kDa | - |
Name: Pre-mRNA-splicing helicase BRR2
EC number: 3.6.4.13
Number of copies: 1
UniProtKB PDBe-KB AlphaFold DB
EC number: 3.6.4.13
Number of copies: 1
UniProtKB PDBe-KB AlphaFold DB
Domains
Gene Ontology [11]
Biological process:
mRNA splicing, via spliceosome spliceosome conformational change to release U4 (or U4atac) and U1 (or U11)
Molecular function:
ATP binding RNA helicase activity identical protein binding nucleic acid binding ATP hydrolysis activity
Cellular location:
U5 snRNP U4/U6 x U5 tri-snRNP complex spliceosomal complex nucleus
mRNA splicing, via spliceosome spliceosome conformational change to release U4 (or U4atac) and U1 (or U11)
Molecular function:
ATP binding RNA helicase activity identical protein binding nucleic acid binding ATP hydrolysis activity
Cellular location:
U5 snRNP U4/U6 x U5 tri-snRNP complex spliceosomal complex nucleus
Natural source
Molecular weight
| Experimental | Theoretical | Method |
|---|---|---|
| - | 246 kDa | - |
Name: Pre-mRNA-splicing factor SNU114
Number of copies: 1
UniProtKB PDBe-KB AlphaFold DB
Number of copies: 1
UniProtKB PDBe-KB AlphaFold DB
Domains
Gene Ontology [17]
Biological process:
spliceosomal snRNP assembly generation of catalytic spliceosome for first transesterification step spliceosomal tri-snRNP complex assembly mRNA splicing, via spliceosome spliceosome conformational change to release U4 (or U4atac) and U1 (or U11)
Molecular function:
translation elongation factor activity GTP binding U5 snRNA binding GTPase activity
Cellular location:
cytosol spliceosomal complex cytoplasm U4/U6 x U5 tri-snRNP complex U2-type catalytic step 2 spliceosome nucleus ribonucleoprotein complex U5 snRNP
spliceosomal snRNP assembly generation of catalytic spliceosome for first transesterification step spliceosomal tri-snRNP complex assembly mRNA splicing, via spliceosome spliceosome conformational change to release U4 (or U4atac) and U1 (or U11)
Molecular function:
translation elongation factor activity GTP binding U5 snRNA binding GTPase activity
Cellular location:
cytosol spliceosomal complex cytoplasm U4/U6 x U5 tri-snRNP complex U2-type catalytic step 2 spliceosome nucleus ribonucleoprotein complex U5 snRNP
Natural source
Molecular weight
| Experimental | Theoretical | Method |
|---|---|---|
| - | 114 kDa | - |
Name: Pre-mRNA-splicing factor RSE1
Number of copies: 1
UniProtKB PDBe-KB AlphaFold DB
Number of copies: 1
UniProtKB PDBe-KB AlphaFold DB
Domains
Gene Ontology [9]
Biological process:
U2-type prespliceosome assembly spliceosomal complex assembly mRNA splicing, via spliceosome
Molecular function:
U2 snRNA binding
Cellular location:
spliceosomal complex nucleus U2 snRNP U2-type spliceosomal complex U2-type prespliceosome
U2-type prespliceosome assembly spliceosomal complex assembly mRNA splicing, via spliceosome
Molecular function:
U2 snRNA binding
Cellular location:
spliceosomal complex nucleus U2 snRNP U2-type spliceosomal complex U2-type prespliceosome
Natural source
Molecular weight
| Experimental | Theoretical | Method |
|---|---|---|
| - | 153 kDa | - |
Name: Cold sensitive U2 snRNA suppressor 1
Number of copies: 1
UniProtKB PDBe-KB AlphaFold DB
Number of copies: 1
UniProtKB PDBe-KB AlphaFold DB
Domains
Gene Ontology [11]
Biological process:
U2-type prespliceosome assembly spliceosomal complex assembly mRNA splicing, via spliceosome
Molecular function:
RNA binding
Cellular location:
spliceosomal complex nucleus U2 snRNP U2-type prespliceosome catalytic step 2 spliceosome precatalytic spliceosome U2-type spliceosomal complex
U2-type prespliceosome assembly spliceosomal complex assembly mRNA splicing, via spliceosome
Molecular function:
RNA binding
Cellular location:
spliceosomal complex nucleus U2 snRNP U2-type prespliceosome catalytic step 2 spliceosome precatalytic spliceosome U2-type spliceosomal complex
Natural source
Molecular weight
| Experimental | Theoretical | Method |
|---|---|---|
| - | 50 kDa | - |
Name: Pre-mRNA-splicing factor PRP11
Number of copies: 1
UniProtKB PDBe-KB AlphaFold DB
Number of copies: 1
UniProtKB PDBe-KB AlphaFold DB
Domains
Gene Ontology [10]
Biological process:
mRNA splicing, via spliceosome U2-type prespliceosome assembly spliceosomal complex assembly
Molecular function:
zinc ion binding RNA binding
Cellular location:
nucleus spliceosomal complex U2-type prespliceosome U2 snRNP catalytic step 2 spliceosome
mRNA splicing, via spliceosome U2-type prespliceosome assembly spliceosomal complex assembly
Molecular function:
zinc ion binding RNA binding
Cellular location:
nucleus spliceosomal complex U2-type prespliceosome U2 snRNP catalytic step 2 spliceosome
Natural source
Molecular weight
| Experimental | Theoretical | Method |
|---|---|---|
| - | 29 kDa | - |
Name: Pre-mRNA-splicing factor RDS3
Number of copies: 1
UniProtKB PDBe-KB AlphaFold DB
Number of copies: 1
UniProtKB PDBe-KB AlphaFold DB
Domains
Gene Ontology [9]
Natural source
Molecular weight
| Experimental | Theoretical | Method |
|---|---|---|
| - | 12 kDa | - |
Name: RDS3 complex subunit 10
Number of copies: 1
UniProtKB PDBe-KB AlphaFold DB
Number of copies: 1
UniProtKB PDBe-KB AlphaFold DB
Domains
Gene Ontology [8]
Biological process:
U2-type prespliceosome assembly mRNA splicing, via spliceosome spliceosomal complex assembly
Cellular location:
nucleus spliceosomal complex U2 snRNP precatalytic spliceosome U2-type spliceosomal complex
U2-type prespliceosome assembly mRNA splicing, via spliceosome spliceosomal complex assembly
Cellular location:
nucleus spliceosomal complex U2 snRNP precatalytic spliceosome U2-type spliceosomal complex
Natural source
Molecular weight
| Experimental | Theoretical | Method |
|---|---|---|
| - | 10 kDa | - |
Name: Pre-mRNA-splicing factor PRP46
Number of copies: 1
UniProtKB PDBe-KB AlphaFold DB
Number of copies: 1
UniProtKB PDBe-KB AlphaFold DB
Domains
Gene Ontology [5]
Biological process:
mRNA splicing, via spliceosome
Cellular location:
Prp19 complex catalytic step 2 spliceosome spliceosomal complex cytoplasm
mRNA splicing, via spliceosome
Cellular location:
Prp19 complex catalytic step 2 spliceosome spliceosomal complex cytoplasm
Natural source
Molecular weight
| Experimental | Theoretical | Method |
|---|---|---|
| - | 50 kDa | - |
Name: Pre-mRNA-processing protein 45
Number of copies: 1
UniProtKB PDBe-KB AlphaFold DB
Number of copies: 1
UniProtKB PDBe-KB AlphaFold DB
Domains
Gene Ontology [5]
Natural source
Molecular weight
| Experimental | Theoretical | Method |
|---|---|---|
| - | 42 kDa | - |
Name: Pre-mRNA-splicing factor SLT11
Number of copies: 1
UniProtKB PDBe-KB AlphaFold DB
Number of copies: 1
UniProtKB PDBe-KB AlphaFold DB
Domains
Gene Ontology [7]
Biological process:
mRNA splicing, via spliceosome
Molecular function:
pre-mRNA binding U6 snRNA binding
Cellular location:
nucleus U2-type catalytic step 2 spliceosome U2-type catalytic step 1 spliceosome Prp19 complex
mRNA splicing, via spliceosome
Molecular function:
pre-mRNA binding U6 snRNA binding
Cellular location:
nucleus U2-type catalytic step 2 spliceosome U2-type catalytic step 1 spliceosome Prp19 complex
Natural source
Molecular weight
| Experimental | Theoretical | Method |
|---|---|---|
| - | 40 kDa | - |
Name: Pre-mRNA-splicing factor CWC2
Number of copies: 1
UniProtKB PDBe-KB AlphaFold DB
Number of copies: 1
UniProtKB PDBe-KB AlphaFold DB
Domains
Gene Ontology [12]
Biological process:
positive regulation of RNA splicing mRNA splicing, via spliceosome spliceosomal snRNP assembly positive regulation of cell cycle mRNA cis splicing, via spliceosome cell cycle
Molecular function:
metal ion binding U6 snRNA binding pre-mRNA binding
Cellular location:
Prp19 complex U2-type catalytic step 1 spliceosome U2-type catalytic step 2 spliceosome
positive regulation of RNA splicing mRNA splicing, via spliceosome spliceosomal snRNP assembly positive regulation of cell cycle mRNA cis splicing, via spliceosome cell cycle
Molecular function:
metal ion binding U6 snRNA binding pre-mRNA binding
Cellular location:
Prp19 complex U2-type catalytic step 1 spliceosome U2-type catalytic step 2 spliceosome
Natural source
Molecular weight
| Experimental | Theoretical | Method |
|---|---|---|
| - | 38 kDa | - |
Name: Pre-mRNA-splicing factor CWC15
Number of copies: 1
UniProtKB PDBe-KB AlphaFold DB
Number of copies: 1
UniProtKB PDBe-KB AlphaFold DB
Domains
Gene Ontology [6]
Biological process:
mRNA splicing, via spliceosome mRNA cis splicing, via spliceosome
Molecular function:
RNA binding
Cellular location:
catalytic step 2 spliceosome U2-type spliceosomal complex nucleus
mRNA splicing, via spliceosome mRNA cis splicing, via spliceosome
Molecular function:
RNA binding
Cellular location:
catalytic step 2 spliceosome U2-type spliceosomal complex nucleus
Natural source
Molecular weight
| Experimental | Theoretical | Method |
|---|---|---|
| - | 19 kDa | - |
Name: Pre-mRNA-splicing factor BUD31
Number of copies: 1
UniProtKB PDBe-KB AlphaFold DB
Number of copies: 1
UniProtKB PDBe-KB AlphaFold DB
Domains
Gene Ontology [5]
Biological process:
cellular bud site selection mRNA splicing, via spliceosome
Cellular location:
nucleus U2 snRNP spliceosomal complex
cellular bud site selection mRNA splicing, via spliceosome
Cellular location:
nucleus U2 snRNP spliceosomal complex
Natural source
Molecular weight
| Experimental | Theoretical | Method |
|---|---|---|
| - | 18 kDa | - |
Name: Pre-mRNA leakage protein 1
Number of copies: 1
UniProtKB PDBe-KB AlphaFold DB
Number of copies: 1
UniProtKB PDBe-KB AlphaFold DB
Domains
Gene Ontology [10]
Biological process:
U2-type prespliceosome assembly maintenance of RNA location mRNA splicing, via spliceosome mRNA export from nucleus
Molecular function:
mRNA binding
Cellular location:
spliceosomal complex U2 snRNP nucleus cytoplasm RES complex
U2-type prespliceosome assembly maintenance of RNA location mRNA splicing, via spliceosome mRNA export from nucleus
Molecular function:
mRNA binding
Cellular location:
spliceosomal complex U2 snRNP nucleus cytoplasm RES complex
Natural source
Molecular weight
| Experimental | Theoretical | Method |
|---|---|---|
| - | 24 kDa | - |
Name: U2 snRNP component IST3
Number of copies: 1
UniProtKB PDBe-KB AlphaFold DB
Number of copies: 1
UniProtKB PDBe-KB AlphaFold DB
Domains
Gene Ontology [13]
Biological process:
U2-type prespliceosome assembly maintenance of RNA location spliceosomal complex assembly mRNA export from nucleus mRNA splicing, via spliceosome generation of catalytic spliceosome for first transesterification step
Molecular function:
RNA binding
Cellular location:
spliceosomal complex nucleus precatalytic spliceosome cytoplasm U2 snRNP RES complex
U2-type prespliceosome assembly maintenance of RNA location spliceosomal complex assembly mRNA export from nucleus mRNA splicing, via spliceosome generation of catalytic spliceosome for first transesterification step
Molecular function:
RNA binding
Cellular location:
spliceosomal complex nucleus precatalytic spliceosome cytoplasm U2 snRNP RES complex
Natural source
Molecular weight
| Experimental | Theoretical | Method |
|---|---|---|
| - | 17 kDa | - |
Name: Pre-mRNA-splicing factor CWC26
Number of copies: 1
UniProtKB PDBe-KB AlphaFold DB
Number of copies: 1
UniProtKB PDBe-KB AlphaFold DB
Domains
Gene Ontology [9]
Biological process:
U2-type prespliceosome assembly maintenance of RNA location mRNA splicing, via spliceosome
Cellular location:
spliceosomal complex U2 snRNP cytoplasm RES complex U2-type spliceosomal complex nucleus
U2-type prespliceosome assembly maintenance of RNA location mRNA splicing, via spliceosome
Cellular location:
spliceosomal complex U2 snRNP cytoplasm RES complex U2-type spliceosomal complex nucleus
Natural source
Molecular weight
| Experimental | Theoretical | Method |
|---|---|---|
| - | 30 kDa | - |
Name: Pre-mRNA-splicing factor ATP-dependent RNA helicase-like protein PRP2
EC number: 3.6.4.13
Number of copies: 1
UniProtKB PDBe-KB AlphaFold DB
EC number: 3.6.4.13
Number of copies: 1
UniProtKB PDBe-KB AlphaFold DB
Domains
Gene Ontology [9]
Biological process:
generation of catalytic spliceosome for first transesterification step snoRNA splicing
Molecular function:
helicase activity ATP hydrolysis activity RNA helicase activity ATP binding RNA binding ATP-dependent activity, acting on RNA
Cellular location:
U2-type catalytic step 1 spliceosome
generation of catalytic spliceosome for first transesterification step snoRNA splicing
Molecular function:
helicase activity ATP hydrolysis activity RNA helicase activity ATP binding RNA binding ATP-dependent activity, acting on RNA
Cellular location:
U2-type catalytic step 1 spliceosome
Natural source
Molecular weight
| Experimental | Theoretical | Method |
|---|---|---|
| - | 99 kDa | - |
Name: Pre-mRNA-splicing factor CWC22
Number of copies: 1
UniProtKB PDBe-KB AlphaFold DB
Number of copies: 1
UniProtKB PDBe-KB AlphaFold DB
Domains
Gene Ontology [5]
Biological process:
mRNA splicing, via spliceosome
Molecular function:
RNA binding
Cellular location:
U2-type spliceosomal complex catalytic step 2 spliceosome cytoplasm
mRNA splicing, via spliceosome
Molecular function:
RNA binding
Cellular location:
U2-type spliceosomal complex catalytic step 2 spliceosome cytoplasm
Natural source
Molecular weight
| Experimental | Theoretical | Method |
|---|---|---|
| - | 67 kDa | - |
Name: Pre-mRNA-splicing factor CWC24
Number of copies: 1
UniProtKB PDBe-KB AlphaFold DB
Number of copies: 1
UniProtKB PDBe-KB AlphaFold DB
Domains
Gene Ontology [8]
Biological process:
snoRNA splicing generation of catalytic spliceosome for first transesterification step mRNA splicing, via spliceosome
Molecular function:
DNA binding metal ion binding
Cellular location:
nucleus spliceosomal complex U2-type spliceosomal complex
snoRNA splicing generation of catalytic spliceosome for first transesterification step mRNA splicing, via spliceosome
Molecular function:
DNA binding metal ion binding
Cellular location:
nucleus spliceosomal complex U2-type spliceosomal complex
Natural source
Molecular weight
| Experimental | Theoretical | Method |
|---|---|---|
| - | 29 kDa | - |
Name: Peptidyl-prolyl isomerase CWC27
EC number: 5.2.1.8
Number of copies: 1
UniProtKB PDBe-KB AlphaFold DB
EC number: 5.2.1.8
Number of copies: 1
UniProtKB PDBe-KB AlphaFold DB
Domains
Gene Ontology [7]
Biological process:
protein peptidyl-prolyl isomerization RNA splicing mRNA processing
Molecular function:
peptidyl-prolyl cis-trans isomerase activity
Cellular location:
catalytic step 2 spliceosome U2-type spliceosomal complex cytoplasm
protein peptidyl-prolyl isomerization RNA splicing mRNA processing
Molecular function:
peptidyl-prolyl cis-trans isomerase activity
Cellular location:
catalytic step 2 spliceosome U2-type spliceosomal complex cytoplasm
Natural source
Molecular weight
| Experimental | Theoretical | Method |
|---|---|---|
| - | 35 kDa | - |
Name: Pre-mRNA-splicing factor CEF1
Number of copies: 1
UniProtKB UniProtKB PDBe-KB PDBe-KB AlphaFold DB AlphaFold DB
Number of copies: 1
UniProtKB UniProtKB PDBe-KB PDBe-KB AlphaFold DB AlphaFold DB
Natural source
Molecular weight
| Experimental | Theoretical | Method |
|---|---|---|
| - | 61 kDa | - |
Name: U2 snRNP component HSH155
Number of copies: 1
UniProtKB PDBe-KB AlphaFold DB
Number of copies: 1
UniProtKB PDBe-KB AlphaFold DB
Domains
Gene Ontology [10]
Biological process:
U2-type prespliceosome assembly spliceosomal complex assembly mRNA splicing, via spliceosome
Molecular function:
mRNA binding
Cellular location:
nucleus spliceosomal complex U2-type spliceosomal complex U2 snRNP U2-type prespliceosome catalytic step 2 spliceosome
U2-type prespliceosome assembly spliceosomal complex assembly mRNA splicing, via spliceosome
Molecular function:
mRNA binding
Cellular location:
nucleus spliceosomal complex U2-type spliceosomal complex U2 snRNP U2-type prespliceosome catalytic step 2 spliceosome
Natural source
Molecular weight
| Experimental | Theoretical | Method |
|---|---|---|
| - | 110 kDa | - |
Name: Pre-mRNA-splicing factor CLF1
Number of copies: 1
UniProtKB UniProtKB PDBe-KB PDBe-KB AlphaFold DB AlphaFold DB
Number of copies: 1
UniProtKB UniProtKB PDBe-KB PDBe-KB AlphaFold DB AlphaFold DB
Natural source
Organism: Saccharomyces cerevisiae S288c
Molecular weight
| Experimental | Theoretical | Method |
|---|---|---|
| - | 68 kDa | - |
Name: Pre-mRNA-splicing factor CWC21
Number of copies: 1
UniProtKB PDBe-KB AlphaFold DB
Number of copies: 1
UniProtKB PDBe-KB AlphaFold DB
Domains
Gene Ontology [3]
Biological process:
mRNA splicing, via spliceosome
Cellular location:
cytoplasm U2-type spliceosomal complex
mRNA splicing, via spliceosome
Cellular location:
cytoplasm U2-type spliceosomal complex
Natural source
Molecular weight
| Experimental | Theoretical | Method |
|---|---|---|
| - | 15 kDa | - |
Name: Pre-mRNA-splicing factor SYF1
Number of copies: 1
UniProtKB PDBe-KB AlphaFold DB
Number of copies: 1
UniProtKB PDBe-KB AlphaFold DB
Natural source
Organism: Saccharomyces cerevisiae S288c
Molecular weight
| Experimental | Theoretical | Method |
|---|---|---|
| - | 78 kDa | - |
Name: Protein HSH49
Number of copies: 1
UniProtKB PDBe-KB AlphaFold DB
Number of copies: 1
UniProtKB PDBe-KB AlphaFold DB
Domains
Gene Ontology [13]
Biological process:
U2-type prespliceosome assembly mRNA splicing, via spliceosome
Molecular function:
mRNA 3'-UTR binding poly(A) binding RNA binding poly(U) RNA binding
Cellular location:
spliceosomal complex U2-type prespliceosome ribonucleoprotein complex U2 snRNP nucleus U2-type spliceosomal complex cytosol
U2-type prespliceosome assembly mRNA splicing, via spliceosome
Molecular function:
mRNA 3'-UTR binding poly(A) binding RNA binding poly(U) RNA binding
Cellular location:
spliceosomal complex U2-type prespliceosome ribonucleoprotein complex U2 snRNP nucleus U2-type spliceosomal complex cytosol
Natural source
Molecular weight
| Experimental | Theoretical | Method |
|---|---|---|
| - | 24 kDa | - |
Name: Pre-mRNA-splicing factor SYF2
Number of copies: 1
UniProtKB PDBe-KB AlphaFold DB
Number of copies: 1
UniProtKB PDBe-KB AlphaFold DB
Domains
Gene Ontology [8]
Natural source
Molecular weight
| Experimental | Theoretical | Method |
|---|---|---|
| - | 24 kDa | - |
Name: Pre-mRNA-processing factor 19
EC number: 6.3.2.-
Number of copies: 4
UniProtKB PDBe-KB AlphaFold DB
EC number: 6.3.2.-
Number of copies: 4
UniProtKB PDBe-KB AlphaFold DB
Domains
Gene Ontology [13]
Biological process:
protein K63-linked ubiquitination generation of catalytic spliceosome for first transesterification step DNA repair mRNA splicing, via spliceosome
Molecular function:
identical protein binding ubiquitin-protein transferase activity ubiquitin protein ligase activity
Cellular location:
nucleus U2-type catalytic step 1 spliceosome mitochondrion Prp19 complex cytoplasm cytosol
protein K63-linked ubiquitination generation of catalytic spliceosome for first transesterification step DNA repair mRNA splicing, via spliceosome
Molecular function:
identical protein binding ubiquitin-protein transferase activity ubiquitin protein ligase activity
Cellular location:
nucleus U2-type catalytic step 1 spliceosome mitochondrion Prp19 complex cytoplasm cytosol
Natural source
Molecular weight
| Experimental | Theoretical | Method |
|---|---|---|
| - | 56 kDa | - |
Name: Pre-mRNA-splicing factor SNT309
Number of copies: 1
UniProtKB PDBe-KB AlphaFold DB
Number of copies: 1
UniProtKB PDBe-KB AlphaFold DB
Gene Ontology [3]
Biological process:
mRNA splicing, via spliceosome
Cellular location:
Prp19 complex U2-type catalytic step 1 spliceosome
mRNA splicing, via spliceosome
Cellular location:
Prp19 complex U2-type catalytic step 1 spliceosome
Natural source
Molecular weight
| Experimental | Theoretical | Method |
|---|---|---|
| - | 20 kDa | - |
Name: Pre-mRNA-processing factor 17
Number of copies: 1
UniProtKB PDBe-KB AlphaFold DB
Number of copies: 1
UniProtKB PDBe-KB AlphaFold DB
Domains
Gene Ontology [8]
Biological process:
mRNA branch site recognition mRNA splicing, via spliceosome generation of catalytic spliceosome for second transesterification step mRNA 3'-splice site recognition
Molecular function:
mRNA binding
Cellular location:
nuclear periphery catalytic step 2 spliceosome spliceosomal complex
mRNA branch site recognition mRNA splicing, via spliceosome generation of catalytic spliceosome for second transesterification step mRNA 3'-splice site recognition
Molecular function:
mRNA binding
Cellular location:
nuclear periphery catalytic step 2 spliceosome spliceosomal complex
Natural source
Molecular weight
| Experimental | Theoretical | Method |
|---|---|---|
| - | 52 kDa | - |
Name: Small nuclear ribonucleoprotein-associated protein B
Number of copies: 1
UniProtKB PDBe-KB AlphaFold DB
Number of copies: 1
UniProtKB PDBe-KB AlphaFold DB
Domains
Gene Ontology [19]
Biological process:
mRNA 5'-splice site recognition U2-type prespliceosome assembly spliceosomal complex assembly spliceosomal snRNP assembly 7-methylguanosine cap hypermethylation mRNA splicing, via spliceosome
Molecular function:
RNA binding
Cellular location:
U4/U6 snRNP spliceosomal complex commitment complex nucleus U4 snRNP catalytic step 2 spliceosome U5 snRNP U4/U6 x U5 tri-snRNP complex U2-type prespliceosome U1 snRNP cytoplasm U2 snRNP
mRNA 5'-splice site recognition U2-type prespliceosome assembly spliceosomal complex assembly spliceosomal snRNP assembly 7-methylguanosine cap hypermethylation mRNA splicing, via spliceosome
Molecular function:
RNA binding
Cellular location:
U4/U6 snRNP spliceosomal complex commitment complex nucleus U4 snRNP catalytic step 2 spliceosome U5 snRNP U4/U6 x U5 tri-snRNP complex U2-type prespliceosome U1 snRNP cytoplasm U2 snRNP
Natural source
Molecular weight
| Experimental | Theoretical | Method |
|---|---|---|
| - | 22 kDa | - |
Name: Small nuclear ribonucleoprotein E
Number of copies: 1
UniProtKB PDBe-KB AlphaFold DB
Number of copies: 1
UniProtKB PDBe-KB AlphaFold DB
Domains
Gene Ontology [21]
Biological process:
mRNA 5'-splice site recognition U2-type prespliceosome assembly spliceosomal complex assembly spliceosomal snRNP assembly mRNA splicing, via spliceosome 7-methylguanosine cap hypermethylation
Molecular function:
splicing factor binding RNA binding
Cellular location:
pICln-Sm protein complex commitment complex nucleus spliceosomal complex U4/U6 snRNP cytoplasm U4/U6 x U5 tri-snRNP complex U2-type prespliceosome U4 snRNP U1 snRNP U2 snRNP precatalytic spliceosome U5 snRNP
mRNA 5'-splice site recognition U2-type prespliceosome assembly spliceosomal complex assembly spliceosomal snRNP assembly mRNA splicing, via spliceosome 7-methylguanosine cap hypermethylation
Molecular function:
splicing factor binding RNA binding
Cellular location:
pICln-Sm protein complex commitment complex nucleus spliceosomal complex U4/U6 snRNP cytoplasm U4/U6 x U5 tri-snRNP complex U2-type prespliceosome U4 snRNP U1 snRNP U2 snRNP precatalytic spliceosome U5 snRNP
Natural source
Molecular weight
| Experimental | Theoretical | Method |
|---|---|---|
| - | 10 kDa | - |
Name: Small nuclear ribonucleoprotein F
Number of copies: 1
UniProtKB PDBe-KB AlphaFold DB
Number of copies: 1
UniProtKB PDBe-KB AlphaFold DB
Domains
Gene Ontology [21]
Biological process:
spliceosomal complex assembly U2-type prespliceosome assembly mRNA 5'-splice site recognition spliceosomal snRNP assembly 7-methylguanosine cap hypermethylation mRNA splicing, via spliceosome
Molecular function:
poly(U) RNA binding RNA binding splicing factor binding
Cellular location:
pICln-Sm protein complex U4/U6 snRNP U4 snRNP nucleus commitment complex spliceosomal complex U4/U6 x U5 tri-snRNP complex cytoplasm U2 snRNP U5 snRNP U1 snRNP catalytic step 2 spliceosome
spliceosomal complex assembly U2-type prespliceosome assembly mRNA 5'-splice site recognition spliceosomal snRNP assembly 7-methylguanosine cap hypermethylation mRNA splicing, via spliceosome
Molecular function:
poly(U) RNA binding RNA binding splicing factor binding
Cellular location:
pICln-Sm protein complex U4/U6 snRNP U4 snRNP nucleus commitment complex spliceosomal complex U4/U6 x U5 tri-snRNP complex cytoplasm U2 snRNP U5 snRNP U1 snRNP catalytic step 2 spliceosome
Natural source
Molecular weight
| Experimental | Theoretical | Method |
|---|---|---|
| - | 9 kDa | - |
Name: Small nuclear ribonucleoprotein G
Number of copies: 1
UniProtKB PDBe-KB AlphaFold DB
Number of copies: 1
UniProtKB PDBe-KB AlphaFold DB
Domains
Gene Ontology [24]
Biological process:
mRNA 5'-splice site recognition U2-type prespliceosome assembly spliceosomal complex assembly 7-methylguanosine cap hypermethylation spliceosomal snRNP assembly mRNA splicing, via spliceosome
Molecular function:
RNA binding
Cellular location:
nucleus commitment complex cytoplasm U4/U6 snRNP spliceosomal complex SMN-Sm protein complex U4/U6 x U5 tri-snRNP complex U4 snRNP U1 snRNP U2-type prespliceosome U5 snRNP U2 snRNP U12-type spliceosomal complex ribonucleoprotein complex spliceosomal tri-snRNP complex catalytic step 2 spliceosome precatalytic spliceosome
mRNA 5'-splice site recognition U2-type prespliceosome assembly spliceosomal complex assembly 7-methylguanosine cap hypermethylation spliceosomal snRNP assembly mRNA splicing, via spliceosome
Molecular function:
RNA binding
Cellular location:
nucleus commitment complex cytoplasm U4/U6 snRNP spliceosomal complex SMN-Sm protein complex U4/U6 x U5 tri-snRNP complex U4 snRNP U1 snRNP U2-type prespliceosome U5 snRNP U2 snRNP U12-type spliceosomal complex ribonucleoprotein complex spliceosomal tri-snRNP complex catalytic step 2 spliceosome precatalytic spliceosome
Natural source
Molecular weight
| Experimental | Theoretical | Method |
|---|---|---|
| - | 8 kDa | - |
Name: Small nuclear ribonucleoprotein Sm D3
Number of copies: 1
UniProtKB PDBe-KB AlphaFold DB
Number of copies: 1
UniProtKB PDBe-KB AlphaFold DB
Domains
Gene Ontology [26]
Biological process:
spliceosomal complex assembly mRNA 5'-splice site recognition U2-type prespliceosome assembly spliceosomal snRNP assembly 7-methylguanosine cap hypermethylation mRNA splicing, via spliceosome
Molecular function:
mRNA binding RNA binding
Cellular location:
U4/U6 snRNP cytoplasm spliceosomal complex nucleus spliceosomal tri-snRNP complex precatalytic spliceosome pICln-Sm protein complex U12-type spliceosomal complex U4 snRNP U2-type prespliceosome U2 snRNP SMN-Sm protein complex U1 snRNP U5 snRNP U4/U6 x U5 tri-snRNP complex catalytic step 2 spliceosome commitment complex cytosol
spliceosomal complex assembly mRNA 5'-splice site recognition U2-type prespliceosome assembly spliceosomal snRNP assembly 7-methylguanosine cap hypermethylation mRNA splicing, via spliceosome
Molecular function:
mRNA binding RNA binding
Cellular location:
U4/U6 snRNP cytoplasm spliceosomal complex nucleus spliceosomal tri-snRNP complex precatalytic spliceosome pICln-Sm protein complex U12-type spliceosomal complex U4 snRNP U2-type prespliceosome U2 snRNP SMN-Sm protein complex U1 snRNP U5 snRNP U4/U6 x U5 tri-snRNP complex catalytic step 2 spliceosome commitment complex cytosol
Natural source
Molecular weight
| Experimental | Theoretical | Method |
|---|---|---|
| - | 11 kDa | - |
Name: Small nuclear ribonucleoprotein Sm D1
Number of copies: 1
UniProtKB PDBe-KB AlphaFold DB
Number of copies: 1
UniProtKB PDBe-KB AlphaFold DB
Domains
Gene Ontology [27]
Biological process:
spliceosomal complex assembly mRNA 5'-splice site recognition U2-type prespliceosome assembly 7-methylguanosine cap hypermethylation spliceosomal snRNP assembly mRNA splicing, via spliceosome
Molecular function:
RNA binding mRNA binding
Cellular location:
spliceosomal complex cytoplasm U4/U6 snRNP spliceosomal tri-snRNP complex commitment complex nucleus U5 snRNP U1 snRNP U4 snRNP U2-type prespliceosome U2 snRNP cytosol precatalytic spliceosome pICln-Sm protein complex U4/U6 x U5 tri-snRNP complex small nuclear ribonucleoprotein complex SMN-Sm protein complex U12-type spliceosomal complex catalytic step 2 spliceosome
spliceosomal complex assembly mRNA 5'-splice site recognition U2-type prespliceosome assembly 7-methylguanosine cap hypermethylation spliceosomal snRNP assembly mRNA splicing, via spliceosome
Molecular function:
RNA binding mRNA binding
Cellular location:
spliceosomal complex cytoplasm U4/U6 snRNP spliceosomal tri-snRNP complex commitment complex nucleus U5 snRNP U1 snRNP U4 snRNP U2-type prespliceosome U2 snRNP cytosol precatalytic spliceosome pICln-Sm protein complex U4/U6 x U5 tri-snRNP complex small nuclear ribonucleoprotein complex SMN-Sm protein complex U12-type spliceosomal complex catalytic step 2 spliceosome
Natural source
Molecular weight
| Experimental | Theoretical | Method |
|---|---|---|
| - | 16 kDa | - |
Name: Small nuclear ribonucleoprotein Sm D2
Number of copies: 1
UniProtKB PDBe-KB AlphaFold DB
Number of copies: 1
UniProtKB PDBe-KB AlphaFold DB
Domains
Gene Ontology [1]
Natural source
Molecular weight
| Experimental | Theoretical | Method |
|---|---|---|
| - | 11 kDa | - |
Name: Saccharomyces cerevisiae strain CDRDR_sf_H chromosome VII sequence
Number of copies: 1
Number of copies: 1
Natural source
Organism: Saccharomyces cerevisiae
Molecular weight
| Experimental | Theoretical | Method |
|---|---|---|
| - | 68 kDa | - |
Name: Saccharomyces cerevisiae strain T.52_2H chromosome XII sequence
Number of copies: 1
Number of copies: 1
Natural source
Organism: Saccharomyces cerevisiae
Molecular weight
| Experimental | Theoretical | Method |
|---|---|---|
| - | 35 kDa | - |
Name: U2 snRNA
Number of copies: 1
Number of copies: 1
Natural source
Organism: Saccharomyces cerevisiae S288c
Molecular weight
| Experimental | Theoretical | Method |
|---|---|---|
| - | 376 kDa | - |
Name: Pre-mRNA
Number of copies: 1
Number of copies: 1
Natural source
Organism: Saccharomyces cerevisiae S288c
Molecular weight
| Experimental | Theoretical | Method |
|---|---|---|
| - | 7 kDa | - |
Name: Pre-mRNA
Number of copies: 1
Number of copies: 1
Natural source
Organism: Saccharomyces cerevisiae S288c
Molecular weight
| Experimental | Theoretical | Method |
|---|---|---|
| - | 18 kDa | - |
Name: GUANOSINE-5'-TRIPHOSPHATE
HET code: GTP
Number of copies: 1
ChEMBL ChEBI DrugBank
HET code: GTP
Number of copies: 1
ChEMBL ChEBI DrugBank
Molecular weight
| Experimental | Theoretical | Method |
|---|---|---|
| - | 523 Da | - |
Name: MAGNESIUM ION
HET code: MG
Number of copies: 5
ChEBI DrugBank
HET code: MG
Number of copies: 5
ChEBI DrugBank
Molecular weight
| Experimental | Theoretical | Method |
|---|---|---|
| - | 24 Da | - |
Name: ZINC ION
HET code: ZN
Number of copies: 13
ChEMBL ChEBI DrugBank
HET code: ZN
Number of copies: 13
ChEMBL ChEBI DrugBank
Molecular weight
| Experimental | Theoretical | Method |
|---|---|---|
| - | 65 Da | - |
Name: ADENOSINE-5'-DIPHOSPHATE
HET code: ADP
Number of copies: 1
ChEMBL ChEBI DrugBank
HET code: ADP
Number of copies: 1
ChEMBL ChEBI DrugBank
Molecular weight
| Experimental | Theoretical | Method |
|---|---|---|
| - | 427 Da | - |
