EMD-3951
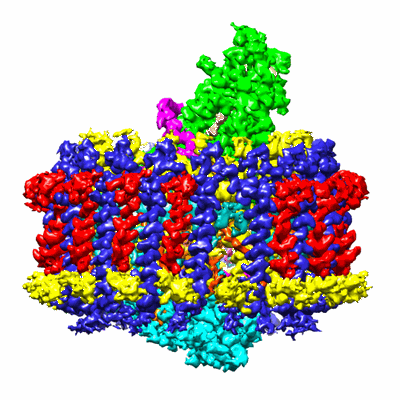
Reaction centre light harvesting complex 1 from Blc. virids
EMD-3951
Single-particle2.87 Å
 Deposition: 25/10/2017
Deposition: 25/10/2017Map released: 11/04/2018
Last modified: 25/11/2020
Name: Reaction centre-Light harvesting complex 1 (RC-LH1)
Summary:
- Complex
- Protein
- Photosynthetic reaction center cytochrome c subunit
- Reaction center protein L chain
- Reaction center protein M chain
- Reaction center protein H chain
- Light-harvesting protein B-1015 alpha chain
- Light-harvesting protein B-1015 beta chain
- Light-harvesting protein B-1015 gamma chain
- Ligand
Name: Reaction centre-Light harvesting complex 1 (RC-LH1)
Details: The protein was isolated from a photosynthetic bacterium Blc. viridis.
Details: The protein was isolated from a photosynthetic bacterium Blc. viridis.
Natural source [1]
Organism: Blastochloris viridis
Molecular weight
| Experimental | Theoretical | Method |
|---|---|---|
| 414 kDa | - | - |
Name: Photosynthetic reaction center cytochrome c subunit
Number of copies: 1
Details: Reaction centre cytochrome subunit
UniProtKB PDBe-KB AlphaFold DB
Number of copies: 1
Details: Reaction centre cytochrome subunit
UniProtKB PDBe-KB AlphaFold DB
Domains
Gene Ontology [7]
Biological process:
photosynthesis, light reaction photosynthesis
Molecular function:
electron transfer activity iron ion binding heme binding
Cellular location:
plasma membrane-derived chromatophore membrane plasma membrane light-harvesting complex
photosynthesis, light reaction photosynthesis
Molecular function:
electron transfer activity iron ion binding heme binding
Cellular location:
plasma membrane-derived chromatophore membrane plasma membrane light-harvesting complex
Natural source
Organism: Rhodopseudomonas viridis
Molecular weight
| Experimental | Theoretical | Method |
|---|---|---|
| - | 37 kDa | - |
Name: Reaction center protein L chain
Number of copies: 1
Details: Reaction centre L subunit
UniProtKB PDBe-KB AlphaFold DB
Number of copies: 1
Details: Reaction centre L subunit
UniProtKB PDBe-KB AlphaFold DB
Domains
Gene Ontology [8]
Biological process:
photosynthetic electron transport in photosystem II protein-chromophore linkage
Molecular function:
metal ion binding bacteriochlorophyll binding electron transporter, transferring electrons within the cyclic electron transport pathway of photosynthesis activity
Cellular location:
plasma membrane-derived chromatophore membrane integral component of membrane plasma membrane light-harvesting complex
photosynthetic electron transport in photosystem II protein-chromophore linkage
Molecular function:
metal ion binding bacteriochlorophyll binding electron transporter, transferring electrons within the cyclic electron transport pathway of photosynthesis activity
Cellular location:
plasma membrane-derived chromatophore membrane integral component of membrane plasma membrane light-harvesting complex
Natural source
Organism: Rhodopseudomonas viridis
Molecular weight
| Experimental | Theoretical | Method |
|---|---|---|
| - | 30 kDa | - |
Name: Reaction center protein M chain
Number of copies: 1
Details: Teaction centre M subunit
UniProtKB PDBe-KB AlphaFold DB
Number of copies: 1
Details: Teaction centre M subunit
UniProtKB PDBe-KB AlphaFold DB
Domains
Gene Ontology [8]
Biological process:
protein-chromophore linkage photosynthetic electron transport in photosystem II
Molecular function:
bacteriochlorophyll binding electron transporter, transferring electrons within the cyclic electron transport pathway of photosynthesis activity metal ion binding
Cellular location:
plasma membrane-derived chromatophore membrane integral component of membrane plasma membrane light-harvesting complex
protein-chromophore linkage photosynthetic electron transport in photosystem II
Molecular function:
bacteriochlorophyll binding electron transporter, transferring electrons within the cyclic electron transport pathway of photosynthesis activity metal ion binding
Cellular location:
plasma membrane-derived chromatophore membrane integral component of membrane plasma membrane light-harvesting complex
Natural source
Organism: Rhodopseudomonas viridis
Molecular weight
| Experimental | Theoretical | Method |
|---|---|---|
| - | 35 kDa | - |
Name: Reaction center protein H chain
Number of copies: 1
Details: Reaction centre H subunit
UniProtKB PDBe-KB AlphaFold DB
Number of copies: 1
Details: Reaction centre H subunit
UniProtKB PDBe-KB AlphaFold DB
Domains
Gene Ontology [7]
Biological process:
protein-chromophore linkage photosynthesis, light reaction
Molecular function:
bacteriochlorophyll binding electron transporter, transferring electrons within the cyclic electron transport pathway of photosynthesis activity
Cellular location:
integral component of membrane plasma membrane-derived chromatophore membrane plasma membrane light-harvesting complex
protein-chromophore linkage photosynthesis, light reaction
Molecular function:
bacteriochlorophyll binding electron transporter, transferring electrons within the cyclic electron transport pathway of photosynthesis activity
Cellular location:
integral component of membrane plasma membrane-derived chromatophore membrane plasma membrane light-harvesting complex
Natural source
Organism: Rhodopseudomonas viridis
Molecular weight
| Experimental | Theoretical | Method |
|---|---|---|
| - | 28 kDa | - |
Name: Light-harvesting protein B-1015 alpha chain
Number of copies: 17
Details: Light harvesting complex 1 alpha subunit
UniProtKB PDBe-KB AlphaFold DB
Number of copies: 17
Details: Light harvesting complex 1 alpha subunit
UniProtKB PDBe-KB AlphaFold DB
Domains
Gene Ontology [5]
Biological process:
photosynthesis, light reaction
Molecular function:
electron transporter, transferring electrons within the cyclic electron transport pathway of photosynthesis activity
Cellular location:
organelle inner membrane integral component of membrane plasma membrane light-harvesting complex
photosynthesis, light reaction
Molecular function:
electron transporter, transferring electrons within the cyclic electron transport pathway of photosynthesis activity
Cellular location:
organelle inner membrane integral component of membrane plasma membrane light-harvesting complex
Natural source
Organism: Rhodopseudomonas viridis
Molecular weight
| Experimental | Theoretical | Method |
|---|---|---|
| - | 6 kDa | - |
Name: Light-harvesting protein B-1015 beta chain
Number of copies: 17
Details: Light harvesting complex 1 beta subunit
UniProtKB PDBe-KB AlphaFold DB
Number of copies: 17
Details: Light harvesting complex 1 beta subunit
UniProtKB PDBe-KB AlphaFold DB
Domains
Gene Ontology [4]
Biological process:
photosynthesis, light reaction
Molecular function:
electron transporter, transferring electrons within the cyclic electron transport pathway of photosynthesis activity
Cellular location:
integral component of membrane plasma membrane light-harvesting complex
photosynthesis, light reaction
Molecular function:
electron transporter, transferring electrons within the cyclic electron transport pathway of photosynthesis activity
Cellular location:
integral component of membrane plasma membrane light-harvesting complex
Natural source
Organism: Rhodopseudomonas viridis
Molecular weight
| Experimental | Theoretical | Method |
|---|---|---|
| - | 6 kDa | - |
Name: Light-harvesting protein B-1015 gamma chain
Number of copies: 16
Details: Light harvesting complex 1 gamma subunit
UniProtKB PDBe-KB AlphaFold DB
Number of copies: 16
Details: Light harvesting complex 1 gamma subunit
UniProtKB PDBe-KB AlphaFold DB
Natural source
Organism: Rhodopseudomonas viridis
Molecular weight
| Experimental | Theoretical | Method |
|---|---|---|
| - | 2 kDa | - |
Name: PROTOPORPHYRIN IX CONTAINING FE
HET code: HEM
Number of copies: 4
ChEBI
HET code: HEM
Number of copies: 4
ChEBI
Molecular weight
| Experimental | Theoretical | Method |
|---|---|---|
| - | 616 Da | - |
Name: BACTERIOCHLOROPHYLL B
HET code: BCB
Number of copies: 38
HET code: BCB
Number of copies: 38
Molecular weight
| Experimental | Theoretical | Method |
|---|---|---|
| - | 909 Da | - |
Name: BACTERIOPHEOPHYTIN B
HET code: BPB
Number of copies: 2
HET code: BPB
Number of copies: 2
Molecular weight
| Experimental | Theoretical | Method |
|---|---|---|
| - | 887 Da | - |
Name: Ubiquinone-9
HET code: UQ9
Number of copies: 2
HET code: UQ9
Number of copies: 2
Molecular weight
| Experimental | Theoretical | Method |
|---|---|---|
| - | 795 Da | - |
Name: FE (III) ION
HET code: FE
Number of copies: 1
ChEBI DrugBank
HET code: FE
Number of copies: 1
ChEBI DrugBank
Molecular weight
| Experimental | Theoretical | Method |
|---|---|---|
| - | 55 Da | - |
Name: SULFATE ION
HET code: SO4
Number of copies: 7
ChEBI DrugBank
HET code: SO4
Number of copies: 7
ChEBI DrugBank
Molecular weight
| Experimental | Theoretical | Method |
|---|---|---|
| - | 96 Da | - |
Name: LAURYL DIMETHYLAMINE-N-OXIDE
HET code: LDA
Number of copies: 2
ChEMBL ChEBI DrugBank
HET code: LDA
Number of copies: 2
ChEMBL ChEBI DrugBank
Molecular weight
| Experimental | Theoretical | Method |
|---|---|---|
| - | 229 Da | - |
Name: MENAQUINONE-9
HET code: MQ9
Number of copies: 1
HET code: MQ9
Number of copies: 1
Molecular weight
| Experimental | Theoretical | Method |
|---|---|---|
| - | 785 Da | - |
Name: 15-cis-1,2-dihydroneurosporene
HET code: NS5
Number of copies: 1
HET code: NS5
Number of copies: 1
Molecular weight
| Experimental | Theoretical | Method |
|---|---|---|
| - | 540 Da | - |
Name: all-trans-1,2-dihydroneurosporene
HET code: NS0
Number of copies: 17
HET code: NS0
Number of copies: 17
Molecular weight
| Experimental | Theoretical | Method |
|---|---|---|
| - | 540 Da | - |
