Assemblies
Assembly Name:
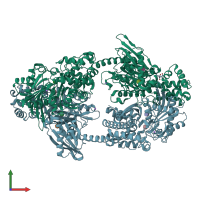
Hexokinase-1
Multimeric state:
homo dimer
Accessible surface area:
69263.26 Å2
Buried surface area:
6217.61 Å2
Dissociation area:
1,807.19
Å2
Dissociation energy (ΔGdiss):
-4.07
kcal/mol
Dissociation entropy (TΔSdiss):
16.71
kcal/mol
Symmetry number:
2
PDBe Complex ID:
PDB-CPX-148564
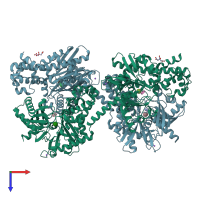
Assembly Name:
Hexokinase-1
Multimeric state:
monomeric
Accessible surface area:
36399.68 Å2
Buried surface area:
1319.07 Å2
Dissociation area:
144.71
Å2
Dissociation energy (ΔGdiss):
-4.48
kcal/mol
Dissociation entropy (TΔSdiss):
2.79
kcal/mol
Symmetry number:
1
PDBe Complex ID:
PDB-CPX-148563
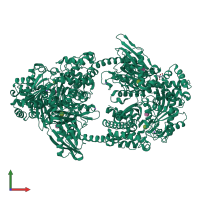
Assembly Name:
Hexokinase-1
Multimeric state:
monomeric
Accessible surface area:
36477.97 Å2
Buried surface area:
1284.15 Å2
Dissociation area:
121.86
Å2
Dissociation energy (ΔGdiss):
-5.18
kcal/mol
Dissociation entropy (TΔSdiss):
2.77
kcal/mol
Symmetry number:
1
PDBe Complex ID:
PDB-CPX-148563
Macromolecules
Chains: A, B
Length: 917 amino acids
Theoretical weight: 102.63 KDa
Source organism: Homo sapiens
Expression system: Escherichia coli
UniProt:
Pfam:
InterPro:
Length: 917 amino acids
Theoretical weight: 102.63 KDa
Source organism: Homo sapiens
Expression system: Escherichia coli
UniProt:
- Canonical:
 P19367 (Residues: 1-917; Coverage: 100%)
P19367 (Residues: 1-917; Coverage: 100%)
Pfam:
InterPro:
- Hexokinase
- ATPase, nucleotide binding domain
- Hexokinase, N-terminal
- Hexokinase, binding site
- Hexokinase, C-terminal