Assemblies
Assembly Name:
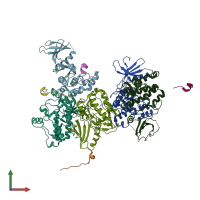
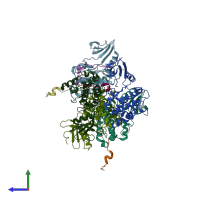
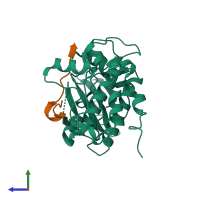
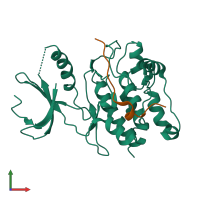
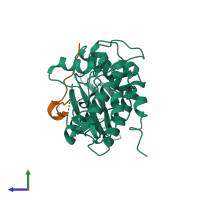
Calcium/calmodulin-dependent protein kinase type II and Calcium/calmodulin-dependent protein kinase II inhibitor 1
Multimeric state:
hetero dimer
Accessible surface area:
13614.11 Å2
Buried surface area:
2038.99 Å2
Dissociation area:
1,019.5
Å2
Dissociation energy (ΔGdiss):
7.07
kcal/mol
Dissociation entropy (TΔSdiss):
8.86
kcal/mol
Symmetry number:
1
PDBe Complex ID:
PDB-CPX-130253
Assembly Name:
Calcium/calmodulin-dependent protein kinase type II and Calcium/calmodulin-dependent protein kinase II inhibitor 1
Multimeric state:
hetero dimer
Accessible surface area:
13275.36 Å2
Buried surface area:
2160.24 Å2
Dissociation area:
1,080.12
Å2
Dissociation energy (ΔGdiss):
7.4
kcal/mol
Dissociation entropy (TΔSdiss):
9.17
kcal/mol
Symmetry number:
1
PDBe Complex ID:
PDB-CPX-130253
Assembly Name:
Calcium/calmodulin-dependent protein kinase type II and Calcium/calmodulin-dependent protein kinase II inhibitor 1
Multimeric state:
hetero dimer
Accessible surface area:
13455.46 Å2
Buried surface area:
2099.03 Å2
Dissociation area:
1,049.51
Å2
Dissociation energy (ΔGdiss):
7.7
kcal/mol
Dissociation entropy (TΔSdiss):
9.06
kcal/mol
Symmetry number:
1
PDBe Complex ID:
PDB-CPX-130253
Assembly Name:
Calcium/calmodulin-dependent protein kinase type II and Calcium/calmodulin-dependent protein kinase II inhibitor 1
Multimeric state:
hetero dimer
Accessible surface area:
13522.73 Å2
Buried surface area:
1966.58 Å2
Dissociation area:
983.29
Å2
Dissociation energy (ΔGdiss):
8.82
kcal/mol
Dissociation entropy (TΔSdiss):
8.98
kcal/mol
Symmetry number:
1
PDBe Complex ID:
PDB-CPX-130253
Assembly Name:
Calcium/calmodulin-dependent protein kinase type II and Calcium/calmodulin-dependent protein kinase II inhibitor 1
Multimeric state:
hetero dimer
Accessible surface area:
13491.6 Å2
Buried surface area:
2119.7 Å2
Dissociation area:
1,059.85
Å2
Dissociation energy (ΔGdiss):
6.85
kcal/mol
Dissociation entropy (TΔSdiss):
9.05
kcal/mol
Symmetry number:
1
PDBe Complex ID:
PDB-CPX-130253
Assembly Name:
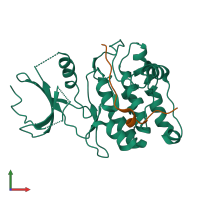
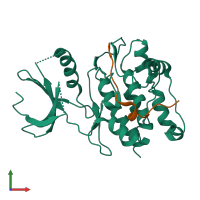
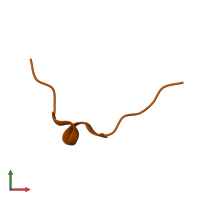
Calcium/calmodulin-dependent protein kinase II inhibitor 1
Multimeric state:
monomeric
Accessible surface area:
2545.13 Å2
Buried surface area:
0.0 Å2
Dissociation area:
0
Å2
Dissociation energy (ΔGdiss):
0
kcal/mol
Dissociation entropy (TΔSdiss):
0
kcal/mol
Symmetry number:
1
PDBe Complex ID:
PDB-CPX-219665
Assembly Name:
Calcium/calmodulin-dependent protein kinase II inhibitor 1
Multimeric state:
monomeric
Accessible surface area:
2358.78 Å2
Buried surface area:
0.0 Å2
Dissociation area:
0
Å2
Dissociation energy (ΔGdiss):
0
kcal/mol
Dissociation entropy (TΔSdiss):
0
kcal/mol
Symmetry number:
1
PDBe Complex ID:
PDB-CPX-219665
Assembly Name:
Calcium/calmodulin-dependent protein kinase II inhibitor 1
Multimeric state:
monomeric
Accessible surface area:
2423.05 Å2
Buried surface area:
0.0 Å2
Dissociation area:
0
Å2
Dissociation energy (ΔGdiss):
0
kcal/mol
Dissociation entropy (TΔSdiss):
0
kcal/mol
Symmetry number:
1
PDBe Complex ID:
PDB-CPX-219665
Macromolecules
Chains: A, C, E, G, I
Length: 269 amino acids
Theoretical weight: 30.45 KDa
Source organism: Caenorhabditis elegans
Expression system: Escherichia coli
UniProt:
Pfam: Protein kinase domain
InterPro:
CATH:
Length: 269 amino acids
Theoretical weight: 30.45 KDa
Source organism: Caenorhabditis elegans
Expression system: Escherichia coli
UniProt:
- Canonical:
 O62305 (Residues: 5-273; Coverage: 37%)
O62305 (Residues: 5-273; Coverage: 37%)
Pfam: Protein kinase domain
InterPro:
CATH:
Chains: B, D, F, H, J
Length: 18 amino acids
Theoretical weight: 2.01 KDa
Source organism: Rattus norvegicus
Expression system: Escherichia coli
UniProt:
InterPro: Calcium/calmodulin-dependent protein kinase II inhibitor
Length: 18 amino acids
Theoretical weight: 2.01 KDa
Source organism: Rattus norvegicus
Expression system: Escherichia coli
UniProt:
- Canonical:
 Q9JI15 (Residues: 42-58; Coverage: 22%)
Q9JI15 (Residues: 42-58; Coverage: 22%)
InterPro: Calcium/calmodulin-dependent protein kinase II inhibitor