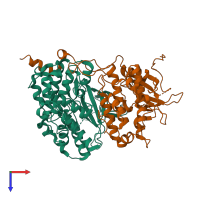
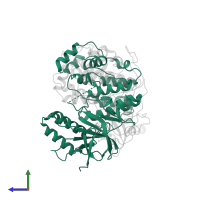
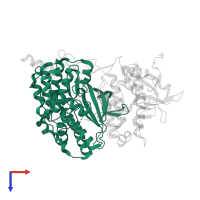
Assemblies
Multimeric state:
hetero dimer
Accessible surface area:
31959.44 Å2
Buried surface area:
3523.69 Å2
Dissociation area:
1,761.85
Å2
Dissociation energy (ΔGdiss):
11.19
kcal/mol
Dissociation entropy (TΔSdiss):
13.89
kcal/mol
Symmetry number:
1
PDBe Complex ID:
PDB-CPX-155866
Multimeric state:
hetero dimer
Accessible surface area:
31865.45 Å2
Buried surface area:
3207.84 Å2
Dissociation area:
1,603.92
Å2
Dissociation energy (ΔGdiss):
2.93
kcal/mol
Dissociation entropy (TΔSdiss):
13.84
kcal/mol
Symmetry number:
1
PDBe Complex ID:
PDB-CPX-155866
Macromolecules
Chains: A, B
Length: 366 amino acids
Theoretical weight: 42 KDa
Source organism: Homo sapiens
Expression system: Spodoptera frugiperda
UniProt:
Pfam: Protein kinase domain
InterPro:
SCOP: Protein kinases, catalytic subunit
Length: 366 amino acids
Theoretical weight: 42 KDa
Source organism: Homo sapiens
Expression system: Spodoptera frugiperda
UniProt:
- Canonical:
 Q16539 (Residues: 2-360; Coverage: 100%)
Q16539 (Residues: 2-360; Coverage: 100%)
Pfam: Protein kinase domain
InterPro:
- Mitogen-activated protein kinase 14
- Protein kinase-like domain superfamily
- Mitogen-activated protein (MAP) kinase p38-like
- Protein kinase domain
- Protein kinase, ATP binding site
- Mitogen-activated protein (MAP) kinase, conserved site
SCOP: Protein kinases, catalytic subunit
Chains: C, D
Length: 406 amino acids
Theoretical weight: 46.29 KDa
Source organism: Homo sapiens
Expression system: Spodoptera frugiperda
UniProt:
Pfam: Protein kinase domain
InterPro:
Length: 406 amino acids
Theoretical weight: 46.29 KDa
Source organism: Homo sapiens
Expression system: Spodoptera frugiperda
UniProt:
- Canonical:
 P49137 (Residues: 1-400; Coverage: 100%)
P49137 (Residues: 1-400; Coverage: 100%)
Pfam: Protein kinase domain
InterPro:
- Protein kinase-like domain superfamily
- Protein kinase domain
- Protein kinase, ATP binding site
- Serine/threonine-protein kinase, active site
- MAP kinase activated protein kinase, C-terminal
- Phosphorylase Kinase; domain 1
- Transferase(Phosphotransferase) domain 1
- MAP kinase activated protein kinase 2