Assemblies
Assembly Name:
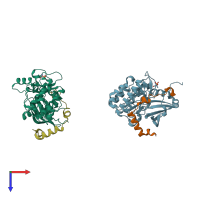
Aurora kinase B-A and Inner centromere protein A
Multimeric state:
hetero dimer
Accessible surface area:
15690.99 Å2
Buried surface area:
3586.2 Å2
Dissociation area:
1,793.1
Å2
Dissociation energy (ΔGdiss):
19.48
kcal/mol
Dissociation entropy (TΔSdiss):
11.14
kcal/mol
Symmetry number:
1
PDBe Complex ID:
PDB-CPX-126740
Assembly Name:
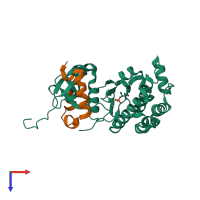
Aurora kinase B-A and Inner centromere protein A
Multimeric state:
hetero dimer
Accessible surface area:
16042.44 Å2
Buried surface area:
3432.18 Å2
Dissociation area:
1,716.09
Å2
Dissociation energy (ΔGdiss):
17.32
kcal/mol
Dissociation entropy (TΔSdiss):
11.01
kcal/mol
Symmetry number:
1
PDBe Complex ID:
PDB-CPX-126740
Macromolecules
Chains: A, B
Length: 284 amino acids
Theoretical weight: 33.54 KDa
Source organism: Xenopus laevis
Expression system: Escherichia coli BL21(DE3)
UniProt:
Pfam: Protein kinase domain
InterPro:
Length: 284 amino acids
Theoretical weight: 33.54 KDa
Source organism: Xenopus laevis
Expression system: Escherichia coli BL21(DE3)
UniProt:
- Canonical:
 Q6DE08 (Residues: 78-361; Coverage: 79%)
Q6DE08 (Residues: 78-361; Coverage: 79%)
Pfam: Protein kinase domain
InterPro:
- Aurora kinase-like
- Protein kinase-like domain superfamily
- Protein kinase domain
- Protein kinase, ATP binding site
- Serine/threonine-protein kinase, active site
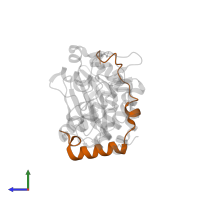
Chains: C, D
Length: 43 amino acids
Theoretical weight: 4.98 KDa
Source organism: Xenopus laevis
Expression system: Escherichia coli BL21(DE3)
UniProt:
Pfam: Inner centromere protein, ARK binding region
InterPro: Inner centromere protein, ARK-binding domain
CATH: Single alpha-helices involved in coiled-coils or other helix-helix interfaces
Length: 43 amino acids
Theoretical weight: 4.98 KDa
Source organism: Xenopus laevis
Expression system: Escherichia coli BL21(DE3)
UniProt:
- Canonical:
 O13024 (Residues: 798-840; Coverage: 5%)
O13024 (Residues: 798-840; Coverage: 5%)
Pfam: Inner centromere protein, ARK binding region
InterPro: Inner centromere protein, ARK-binding domain
CATH: Single alpha-helices involved in coiled-coils or other helix-helix interfaces