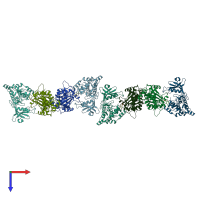
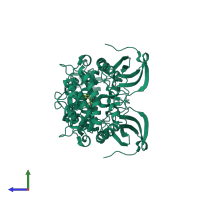
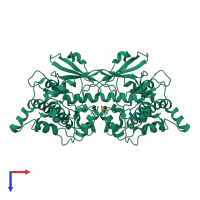
Assemblies
Assembly Name:
Death-associated protein kinase 2
Multimeric state:
homo dimer
Accessible surface area:
27897.98 Å2
Buried surface area:
3694.73 Å2
Dissociation area:
1,563.92
Å2
Dissociation energy (ΔGdiss):
-6.74
kcal/mol
Dissociation entropy (TΔSdiss):
13.79
kcal/mol
Symmetry number:
2
PDBe Complex ID:
PDB-CPX-193792
Assembly Name:
Death-associated protein kinase 2
Multimeric state:
homo dimer
Accessible surface area:
27743.76 Å2
Buried surface area:
3638.15 Å2
Dissociation area:
1,535.45
Å2
Dissociation energy (ΔGdiss):
-8.21
kcal/mol
Dissociation entropy (TΔSdiss):
13.78
kcal/mol
Symmetry number:
2
PDBe Complex ID:
PDB-CPX-193792
Assembly Name:
Death-associated protein kinase 2
Multimeric state:
homo dimer
Accessible surface area:
27772.75 Å2
Buried surface area:
3785.8 Å2
Dissociation area:
1,606.65
Å2
Dissociation energy (ΔGdiss):
-5.29
kcal/mol
Dissociation entropy (TΔSdiss):
13.8
kcal/mol
Symmetry number:
2
PDBe Complex ID:
PDB-CPX-193792
Assembly Name:
Death-associated protein kinase 2
Multimeric state:
homo dimer
Accessible surface area:
27698.63 Å2
Buried surface area:
3878.72 Å2
Dissociation area:
1,657.15
Å2
Dissociation energy (ΔGdiss):
-7.61
kcal/mol
Dissociation entropy (TΔSdiss):
13.81
kcal/mol
Symmetry number:
2
PDBe Complex ID:
PDB-CPX-193792
Macromolecules
Chains: A, B, C, D, E, F, G, H
Length: 321 amino acids
Theoretical weight: 37.03 KDa
Source organism: Homo sapiens
Expression system: Escherichia coli
UniProt:
Pfam: Protein kinase domain
InterPro:
Length: 321 amino acids
Theoretical weight: 37.03 KDa
Source organism: Homo sapiens
Expression system: Escherichia coli
UniProt:
- Canonical:
 Q9UIK4 (Residues: 11-330; Coverage: 87%)
Q9UIK4 (Residues: 11-330; Coverage: 87%)
Pfam: Protein kinase domain
InterPro:
- Protein kinase-like domain superfamily
- Protein kinase domain
- Protein kinase, ATP binding site
- Serine/threonine-protein kinase, active site