Assemblies
Assembly Name:
Halotolerant alpha-type carbonic anhydrase (dca ii)
Multimeric state:
monomeric
Accessible surface area:
11831.05 Å2
Buried surface area:
222.74 Å2
Dissociation area:
61.37
Å2
Dissociation energy (ΔGdiss):
10.4
kcal/mol
Dissociation entropy (TΔSdiss):
-1.11
kcal/mol
Symmetry number:
1
PDBe Complex ID:
PDB-CPX-162560
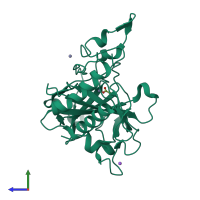
Assembly Name:
Halotolerant alpha-type carbonic anhydrase (dca ii)
Multimeric state:
monomeric
Accessible surface area:
11751.34 Å2
Buried surface area:
303.64 Å2
Dissociation area:
61.6
Å2
Dissociation energy (ΔGdiss):
10.39
kcal/mol
Dissociation entropy (TΔSdiss):
-1.11
kcal/mol
Symmetry number:
1
PDBe Complex ID:
PDB-CPX-162560
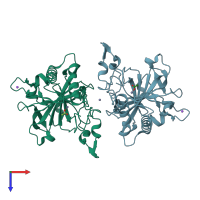
Assembly Name:
Halotolerant alpha-type carbonic anhydrase (dca ii)
Multimeric state:
homo dimer
Accessible surface area:
22363.0 Å2
Buried surface area:
1745.77 Å2
Dissociation area:
650.14
Å2
Dissociation energy (ΔGdiss):
35.27
kcal/mol
Dissociation entropy (TΔSdiss):
12.92
kcal/mol
Symmetry number:
2
PDBe Complex ID:
PDB-CPX-162561
Macromolecules
Chains: A, B
Length: 291 amino acids
Theoretical weight: 31.7 KDa
Source organism: Dunaliella salina
Expression system: Escherichia coli
Length: 291 amino acids
Theoretical weight: 31.7 KDa
Source organism: Dunaliella salina
Expression system: Escherichia coli
- Alpha carbonic anhydrase domain superfamily
- Alpha carbonic anhydrase domain
- Carbonic anhydrase, prokaryotic-like
- Carbonic anhydrase, alpha-class
- Carbonic anhydrase, alpha-class, conserved site