Assemblies
Assembly Name:
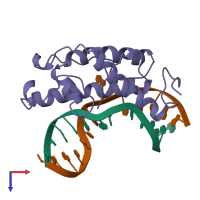
Endonuclease V and DNA
Multimeric state:
hetero trimer
Accessible surface area:
10367.6 Å2
Buried surface area:
3861.38 Å2
Dissociation area:
1,385.48
Å2
Dissociation energy (ΔGdiss):
7.65
kcal/mol
Dissociation entropy (TΔSdiss):
11.18
kcal/mol
Symmetry number:
1
PDBe Complex ID:
PDB-CPX-114719
Macromolecules
Chain: A
Length: 137 amino acids
Theoretical weight: 15.97 KDa
Source organism: Escherichia virus T4
Expression system: Escherichia coli
UniProt:
InterPro:
SCOP: T4 endonuclease V
Length: 137 amino acids
Theoretical weight: 15.97 KDa
Source organism: Escherichia virus T4
Expression system: Escherichia coli
UniProt:
- Canonical:
 P04418 (Residues: 2-138; Coverage: 99%)
P04418 (Residues: 2-138; Coverage: 99%)
InterPro:
- Pyrimidine dimer DNA glycosylase
- Pyrimidine dimer DNA glycosylase, endonuclease V type
- T4 endonuclease V
SCOP: T4 endonuclease V
Name:
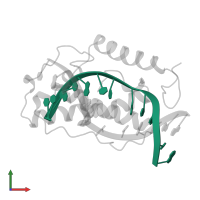
DNA (5'-D(*AP*TP*CP*GP*CP*GP*TP*TP*GP*CP*GP*CP*T)-3')
Representative chains: B
Source organism: Tequatrovirus T4 [10665]
Expression system: Not provided
Length: 13 nucleotides
Theoretical weight: 3.96 KDa
Representative chains: B
Source organism: Tequatrovirus T4 [10665]
Expression system: Not provided
Length: 13 nucleotides
Theoretical weight: 3.96 KDa
Name:
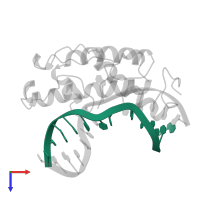
DNA (5'-D(*TP*AP*GP*CP*GP*CP*AP*AP*CP*GP*CP*GP*A)-3')
Representative chains: C
Source organism: Tequatrovirus T4 [10665]
Expression system: Not provided
Length: 13 nucleotides
Theoretical weight: 3.99 KDa
Representative chains: C
Source organism: Tequatrovirus T4 [10665]
Expression system: Not provided
Length: 13 nucleotides
Theoretical weight: 3.99 KDa