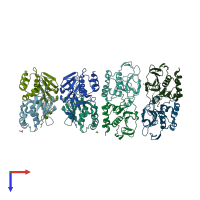Assemblies
Assembly Name:
Leader protease
Multimeric state:
homo dimer
Accessible surface area:
15138.77 Å2
Buried surface area:
3162.87 Å2
Dissociation area:
1,510.44
Å2
Dissociation energy (ΔGdiss):
7.13
kcal/mol
Dissociation entropy (TΔSdiss):
12.05
kcal/mol
Symmetry number:
1
PDBe Complex ID:
PDB-CPX-136753
Assembly Name:
Leader protease
Multimeric state:
homo dimer
Accessible surface area:
15143.04 Å2
Buried surface area:
3388.57 Å2
Dissociation area:
1,512.62
Å2
Dissociation energy (ΔGdiss):
7.26
kcal/mol
Dissociation entropy (TΔSdiss):
12.06
kcal/mol
Symmetry number:
1
PDBe Complex ID:
PDB-CPX-136753
Assembly Name:
Leader protease
Multimeric state:
homo dimer
Accessible surface area:
15187.61 Å2
Buried surface area:
3079.84 Å2
Dissociation area:
1,465.86
Å2
Dissociation energy (ΔGdiss):
7.22
kcal/mol
Dissociation entropy (TΔSdiss):
12.04
kcal/mol
Symmetry number:
1
PDBe Complex ID:
PDB-CPX-136753
Assembly Name:
Leader protease
Multimeric state:
homo dimer
Accessible surface area:
15103.31 Å2
Buried surface area:
2946.59 Å2
Dissociation area:
1,417.52
Å2
Dissociation energy (ΔGdiss):
8.76
kcal/mol
Dissociation entropy (TΔSdiss):
12.03
kcal/mol
Symmetry number:
1
PDBe Complex ID:
PDB-CPX-136753
Macromolecules
Chains: A, B, C, D, E, F, G, H
Length: 173 amino acids
Theoretical weight: 19.78 KDa
Source organism: Foot-and-mouth disease virus (strain O1)
Expression system: Escherichia coli BL21(DE3)
UniProt:
InterPro:
CATH: Cysteine proteinases
SCOP: FMDV leader protease
Length: 173 amino acids
Theoretical weight: 19.78 KDa
Source organism: Foot-and-mouth disease virus (strain O1)
Expression system: Escherichia coli BL21(DE3)
UniProt:
- Canonical:
 P03305 (Residues: 29-201; Coverage: 7%)
P03305 (Residues: 29-201; Coverage: 7%)
InterPro:
CATH: Cysteine proteinases
SCOP: FMDV leader protease