Assemblies
Assembly Name:
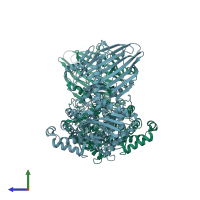
Hydrolase
Multimeric state:
homo dimer
Accessible surface area:
52820.98 Å2
Buried surface area:
4313.44 Å2
Dissociation area:
2,023.48
Å2
Dissociation energy (ΔGdiss):
-4.38
kcal/mol
Dissociation entropy (TΔSdiss):
15.38
kcal/mol
Symmetry number:
1
PDBe Complex ID:
PDB-CPX-181964
Macromolecules
Chains: A, B
Length: 754 amino acids
Theoretical weight: 85.83 KDa
Source organism: Levilactobacillus brevis
UniProt:
Length: 754 amino acids
Theoretical weight: 85.83 KDa
Source organism: Levilactobacillus brevis
UniProt:
- Canonical:
 Q7SIE1 (Residues: 1-754; Coverage: 100%)
Q7SIE1 (Residues: 1-754; Coverage: 100%)
- Glycosyl hydrolase family 65, N-terminal domain
- Glycosyl hydrolase family 65 central catalytic domain
- Glycosyl hydrolase family 65, C-terminal domain
- Galactose mutarotase-like domain superfamily
- Maltose phosphorylase/glycosyl hydrolase/vacuolar acid trehalase
- Glycoside hydrolase family 65, N-terminal domain superfamily
- Glycoside hydrolase, family 65, N-terminal
- Six-hairpin glycosidase superfamily
- Six-hairpin glycosidase-like superfamily
- Glycoside hydrolase, family 65, central catalytic
- Glycoside hydrolase family 65, C-terminal
- Glycoside hydrolase, family 65, N-terminal domain
- Maltose phosphorylase, domain 3
- Glycosyltransferase