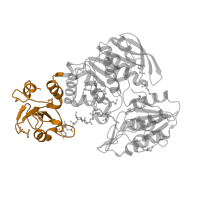
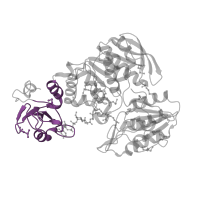
EC 6.3.2.13: UDP-N-acetylmuramoyl-L-alanyl-D-glutamate--2,6-diaminopimelate ligase
Reaction catalysed:
ATP + UDP-N-acetyl-alpha-D-muramoyl-L-alanyl-D-glutamate + meso-2,6-diaminoheptanedioate = ADP + phosphate + UDP-N-acetyl-alpha-D-muramoyl-L-alanyl-D-gamma-glutamyl-meso-2,6-diaminoheptanedioate
Systematic name:
UDP-N-acetyl-alpha-D-muramoyl-L-alanyl-D-glutamate:meso-2,6-diaminoheptanedioate gamma-ligase (ADP-forming)
Alternative Name(s):
- MurE synthetase
- UDP-N-acetylmuramoyl-L-alanyl-D-glutamate:meso-2,6-diamino-heptanedioate ligase (ADP-forming)
- UDP-N-acetylmuramoyl-L-alanyl-D-glutamate:meso-2,6-diaminoheptanedioate gamma-ligase (ADP-forming)
- UDP-N-acetylmuramoyl-L-alanyl-D-glutamyl-meso-2,6-diaminopimelate synthetase
- UDP-N-acetylmuramoylalanyl-D-glutamate--2,6-diaminopimelate ligase
- UDP-N-acetylmuramyl-tripeptide synthetase
GO terms
Biochemical function:
Biological process:
Cellular component:
Sequence families
 Protein families (Pfam)
Protein families (Pfam)
 InterPro annotations
InterPro annotations
IPR005761
Domain description:
UDP-N-acetylmuramoylalanyl-D-glutamate-2,6-diaminopimelate ligase
Occurring in:
Occurring in:
Structure domains
 CATH domains
CATH domains
3.90.190.20
Class:
Alpha Beta
Architecture:
Alpha-Beta Complex
Topology:
Protein-Tyrosine Phosphatase; Chain A
Homology:
Mur ligase, C-terminal domain
Occurring in:

3.40.1190.10
Class:
Alpha Beta
Architecture:
3-Layer(aba) Sandwich
Topology:
UDP-N-acetylmuramoyl-L-alanine:D-glutamate ligase
Homology:
Mur-like, catalytic domain
Occurring in:

 SCOP annotations
SCOP annotations
53245
Class:
Alpha and beta proteins (a/b)
Fold:
MurD-like peptide ligases, peptide-binding domain
Superfamily:
MurD-like peptide ligases, peptide-binding domain
Occurring in:

53624
Class:
Alpha and beta proteins (a/b)
Fold:
Ribokinase-like
Superfamily:
MurD-like peptide ligases, catalytic domain
Occurring in:
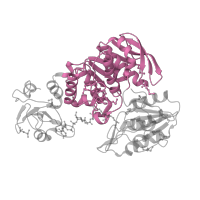
63419
Class:
Alpha and beta proteins (a/b)
Fold:
MurF and HprK N-domain-like
Superfamily:
MurE/MurF N-terminal domain
Occurring in: